Methods for Amplification of PHANTOM DNA
Methods for Amplification of PHANTOM DNA of 547bp with
Polymerase Chain Reaction (PCR) Using Modulated Secondary Broadband
Electromagnetic Radiation (MBER) of LGN-303 Laser
as a Quantum Equivalent of Analogous Material DNA
I am writing about DNA phantoms and the “DNA Phantom Effect”, discovered by myself in 1984
using the method of correlation laser spectroscopy. The first publication about this was in 1991, [Gariaev et
al, 1991, Holographic Associative Memory of Biological Systems, Proceedings SPIE — The International Society for
Optical Engineering. Optical Memory and Neural Networks., v.1621, p.280- 291. USA]. Read more about the
“DNA Phantom Effect” and DNA phantoms in their variety of forms in my monographs “Wave
Genome”'(1994) and “Linguistics Wave Genome. Theory and Practice” (2009). You can find these monographs
with Google.
In Canada (Toronto) in 2001, we used DNA phantoms in the form of secondary radiation from a
LGN-303 laser (http://www.plasmalabs.com/files/products/lgn_303_1.pdf) for the transmission of a pool of
active genetic information over a distance of 20 km. In this experiment, pancreases of a few dozen Wistar
rats were inactivated through alloxan induced diabetes. These rats were subsequently ‘injected’ with distant
phantom (quantum) genetic information, read by the laser from newborn Wistar rats pancreases of the
same genetic lineage. In the control group (not receiving phantom genetic information) 90% of these rats
died. However, all rats that received phantom genetic information, survived. [Gariaev P.P., Kokaya A.A.,
Mukhina I.V., Leonova-Gariaeva E.A., Kokaya N.G. 2007. The effect of biostructures modulated electromagnetic
radiation on alloxan diabetes in rats. Bulletin of Experimental Biology and Medicine, № 2, pp.155-158]. Later, these
findings were confirmed by the independent group of A. Kokaya and the thesis of N. Kokaya, and the group
of G.G. Tertyshniy. These works have been published.
In 2014, we obtained preliminary data on materialization of DNA phantoms, obtained in the form
of spectra of secondary LGN-303 laser radiation, generated by laser reading of DNA segments of a certain
length [Gariaev et al, DNA Decipher Journal / May 2014 Vol. 4 Issue 1/ pp.01-02 Materialization of DNA
Fragment in Water through Modulated Electromagnetic Irradiation. Intern. Patent 2014/06578 08 Sept., 2014].
These experimental results have significant potential value. They confirm A.G. Gurvich’s (1920’s —
40’s) fundamental idea about genes functioning as wave forms. This means that now we can work in the
field of new genetics and, consequently, new biology, medicine, agriculture and computing, and have a new
approach to the problem of the origin of life on Earth, and so on. Nevertheless, phantom DNA data as
quantum equivalents of ordinary material DNA, requires extensive reproduction by independent research
groups.
That is why I am making a proposal to molecular biologists and geneticists with an invitation to
take part in a large-scale open experiment for the extensive reproduction of our results. Following is a full
description for the method of detection and materialization of DNA phantoms in a PCR system.
Step 1. Preparation of the Initial DNA Product
The PCR product, 547bp in length, derived from a synthetic DNA sequence, cloned
in a plasmid, was used as the initial DNA product.
The plus DNA strand:

5′-
CCTTACGTCAGTGGAGATGTCACATCAATCAACTTGCTTTGAAGACGTGGTT
GGAACGTCTTCTTTTTCCACGATGCTCCTCGTGGGTGGGGGTCCATCTTTGG
GACCACTGTCGGCAGAGGCATCTTGAATGATAGCCTTTCCTTTATCGCAATGA
TGGCATTTGTAGGAGCCACCTTCCTTTTCTACTGTCCTTGCGCGCTATATTTT
GTTTTCTATCGCGTATTAAATGTATAATTGGGGGACTCTAATCATAAAAACCC
ATCTCATAAATAACGTCATGCATTACATGTTAATTATTACATGCTTAACGTAAT
TCAACAGAAATTATATGATAATCATCGCAAGACCGGCAACAGGATTCAATCTT
AAGAAACTTTATTGCACGCATTAATGGACTGGATTGGGGCCAACTCCTACCG
TACCTGGCATTACCCTTACGCTGAAGAGATGCTCGACTGGGCAGATGAACAT
GGCATCGTGGTGATTGATGAAACTGCTGCTGTCGGCTTTAACCTCTCTTTAG
GCATTGGTTTGGAAGCGGGCA-3′
For DNA production through PCR amplification the following primer pair was used:
5′-CCTTACGTCAGTGGAGATGTCACATC-3′;
5′-TGCCCGCTTCCAAACCAATGCCTAAAGA-3′.
Each PCR mixture with a final volume of 25μL contained: 67mM Tris-HCl pH 8.6 at
25°C; 2.5mM magnesium chloride; 16.6mM ammonium sulfate; dNTPs mix at a total
concentration of 300μM; primer mix in 0.5μM each; 2.5 units of Taq DNA polymerase and
plasmid DNA-template in amount of 25ng. PCR temperature regime included:
- initial denaturation at 94°C — 3 min.;
- 30 cycles of: 94°C — 30 seconds, 62°C — 30 seconds, 72°C — 40 sec;
- final synthesis 72°C for 5 minutes.
The PCR product was purified from primers and other components of the PCR
reaction solution using a set of reagents for purification implementing SiO2 coated
magnetic particles («Sileks», http://www.sileks.com) according to the manufacturer’s
recommendations. 10μL of magnetic particles with binding capacity for 10 mg DNA were
used. The elution of DNA was performed in 50μL of elution buffer.
Step 2. Preparation: Modulated Broadband Electromagnetic Spectrum of
DNA Product.
25μL of the aqueous solution of the PCR product was applied to a clean microscope
slide and used for probing by the helium-neon LGN-303 laser beam for 3 and more
minutes. The resulting secondary modulated broadband electromagnetic radiation was
recorded by a transistor radio at a frequency of 700 kHz and then converted into Waveform
audio file format. This is the WAVE audio file we propose to use for DNA information
induction (without the use of in this version the LGN-303 laser) into samples of purified
water, considering that sound can carry torsion information
http://www.efir.com.ua/rus/a.php?r=2&d=69, including information about DNA (a hypothesis).
In addition, and at the same time the audio recording was made, DNA information
induction into purified distilled water was facilitated by way of a stationary tripod with test
tubes containing purified distilled water without DNA impurities, RNA and nucleases being
placed at a distance of 15-20 cm from the laser. The water was pre-frozen at -20°C and
thawed at room temperature (defrost water).
Step 3. PCR Amplification Using Water Samples Treated by
the Modulated Broadband Electromagnetic Spectrum of the Initial DNA from
which Information was Read by a LGN-303 Laser
After laser exposure, water treated by the modulated broadband electromagnetic
spectrum, was used for execution of standard PCR reactions, 25μL of this water was used
without addition of DNA-template material. Tripod with the tubes was placed at a distance
of 20-30 cm from the laser. Preparation of the PCR reactions was performed in a sterile
environment treated with UV light that would prevent contamination of samples.
Each PCR mixture contained: 67mM Tris-HCl pH 8.6 at 25°C; 2.5mM magnesium
chloride; 16.6mM ammonium sulfate; dNTPs mix at a total concentration of 300μM;
primer mix in 0.5μM each; 2.5 units of Taq DNA polymerase. Several temperature regimes
were used for PCR, different in duration of the elongation stage (synthesis) of DNA strands
at 72°C.
Initial PCR temperature regime included: - initial denaturation at 94°C — 3 min.;
- 40 cycles: 94°C — 30 seconds, 62°C — 30 seconds, 72°C — 40 sec;
- final synthesis at 72°C for 5 minutes.
Modified PCR temperature regime included: - initial denaturation at 94°C — 3 min.;
- 40 cycles: 94°C — 30 seconds, 62°C — 30 seconds, 72°C — 2.7min;
- final synthesis at 72°C for 5-7 minutes.
All temperature regimes resulted in synthesis of PCR products of predetermined
length, but to varying degrees. The largest number of test specimens with the desired
product was obtained when using a seven-minute DNA strand elongation step in each of
the 40 PCR cycles.
Step 4. Analysis of PCR Results
After the completion of the PCR reaction, the samples were mixed with gel loading
buffer, containing a fluorescent SYBR Green I dye («Sileks», http://www.sileks.com) and were
analyzed on a 1.5% agarose gel by standard methods of gel electrophoresis in a single TBE
buffer. The results were analyzed with a transilluminator at a wavelength of 365nm.
Samples were considered positive where bands were located at the same distance from the
start as the positive control strip, corresponding to a known DNA fragment size of 547bp
PCR positive control samples, samples of the initial DNA product from which the
laser reading was made, and the experimental samples were subjected to sequencing.
Sequencing revealed that the experimental samples are 99.2-100% identical to the DNA
sequence of the initial product from which the information was read by the laser.
To illustrate our PCR experiments, the images of the PCR DNA phantoms are
shown. A link for the download of the audio recording of DNA modulated broadband
electromagnetic radiation in WAVE format on a carrier frequency of 700kGts is also
provided.
PCR Exp
21.01.2016.doc
PCR Exp
03.02.2016.doc
A link to download the audio recording of DNA in wave format:
547bp Sound http://files.mail.ru/A59F39CE29C04CD180132D8885580905
Our experiments can be reproduced without the use of a laser and without the original DNAtemplate
(as we do too), by using the audio recording. i.e. no need to synthesize the template. This
prevents accidental drifts of initial DNA into the working tubes with water, the pipettes, etc. Thus,
we avoid contamination. In this case, the working sound player to be set at a distance of 1 to 20 cm
from the tripod with the test tubes. The exposure time may vary from 5 minutes or more. The
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16
identity of the resulting PCR DNA product and initial DNA can be confirmed by sequencing the
obtained DNA. In this type verification, all that is required is the primers and knowledge of the
initial DNA nucleotide sequence…
Gariaev P.P.
Ph.D. Biology, Acad. of Russian Academy of Medical and Technical Sciences, Acad. of Academy
of Natural Sciences,
Acad. of International Academy of Ecology and Life Safety,
Member of the New York Academy of Sciences, Scientific
Director of Institute of Quantum Genetics
DNA Product of 547bp
Studying of the Impact of Pre-audio and the Audio Recording
of MBER Spectra of the Laser (WAVE File), Recorded on 01.12.2015
Step 1. A study of the laser recording background before exposure to the sound WAVE recording
as a derivative of DNA MBER spectrum
Time of PCR cycle 03/02/2016: from 13.10-20.40 (PCR program with 7-minute elongation (synthesis) of
DNA strands)
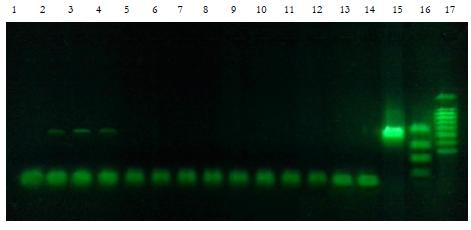
Electrophoresis from 05.02.2016
Tracks 1-14. — The negative control (background before exposure to the sound) — purified water,
commonly used in PCR reactions as the solvent of reaction components, pre-frozen and thawed. The
origin of the water is the same as the water used at the time of laser recording, it was frozen in the
laboratory.
Track 15. — A positive PCR control (plasmid DNA-matrix 25ng, 30 cycles of PCR)
Track 16. — The marker fragments length 300, 400, 500, 600, 700, 800, 900, 1000, 1500bp
 Предыдущий пост
Предыдущий пост Следующий пост
Следующий пост